|
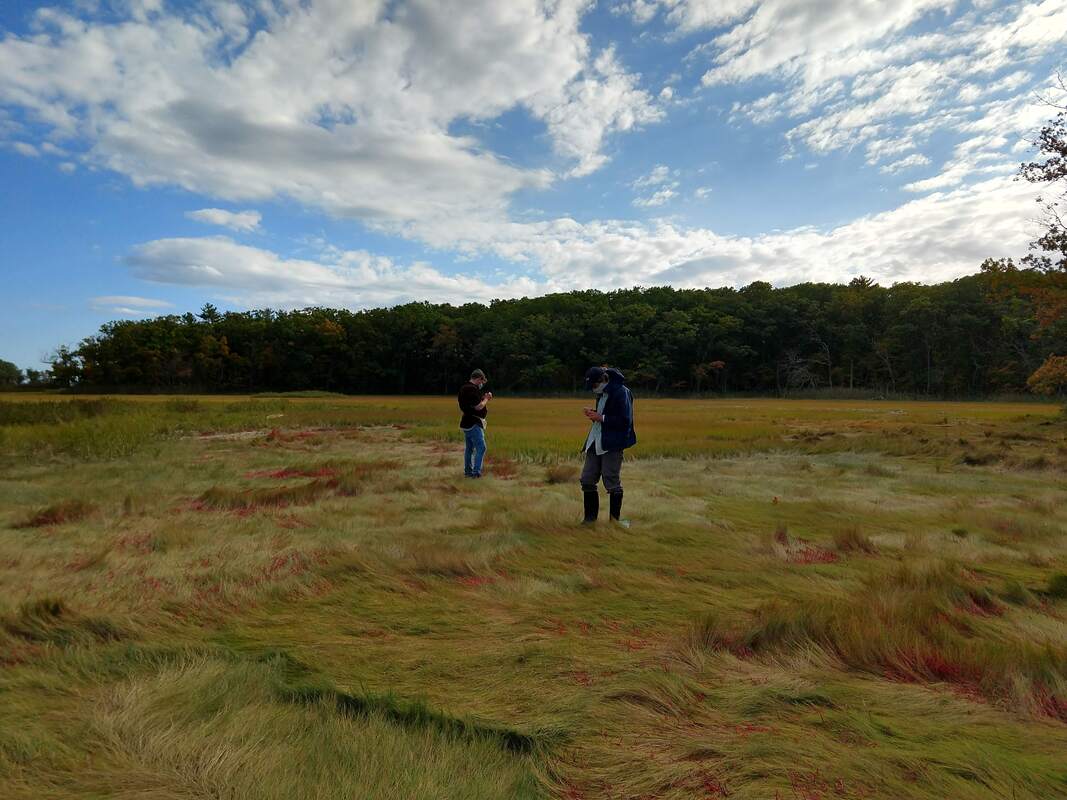
During some recent fieldwork in Manchester, MA, Alice and Cameron were swarmed by the paparazzi (okay, just one local reporter). Their conversation led to a feature on our on-going Salicornia project: https://www.thecricket.com/news/for-the-love-of-salicornia/article_befe49f0-fc41-11eb-9170-77622931e857.html
Alice's hard work has also resulted in a National Science Foundation Graduate Research Fellowship and was highlighted as part of the virtual Evolution 2021 meeting! You can learn more about it by watching her seminar hosted by the Nantucket Field Station last year, on youtube here.
0 Comments
Today was UMass Boston's undergraduate summer research symposium, and the final day of Tadeo Zuniga's REU program here. We had an action-packed summer with a project that combined aspects of engineering, plant biology, and computer science, and I look forward to seeing where Tadeo's career takes him. If you'd like to know more about the project, click through for an abstract.
My first In Brief is online! This format is a way to highlight new and fascinating papers from The Plant Cell, where I serve as an Assistant Features Editor. I take a look at complex epistasis in transcription control of aliphatic glucosinolate biosynthesis (aka that delicious mustard flavor) reported by Li and colleagues (2018). Check it out!
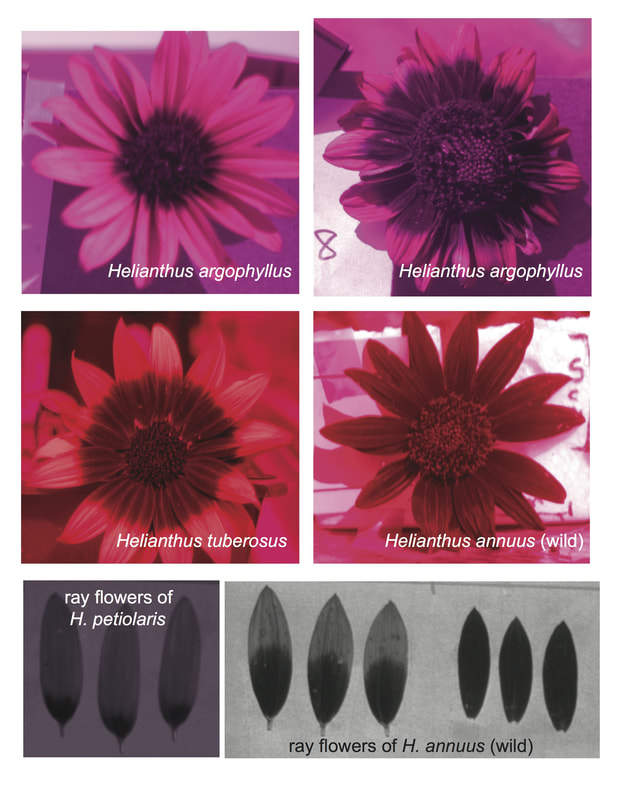
I'm thrilled that our recent paper on ultraviolet-absorbing floral patterns in sunflowers got a write up at Botany One! Read about it at Not For Your Eyes.* * One quick note, what is reported as the "intensity of the pattern" was actually the proportional size of the UV bullseye (how much of the total floral surface it covers), which is controlled by a different part of the genome than the absolute size of the bullseye (its diameter). So, a small flowerhead can be 90% UV absorptive but still have a smaller diameter bullseye than a huge flowerhead that is 50% UV absorbing pigment.
This summer I had the chance to try out teaching the basics of data management and the programming language R to undergraduates in a five week short course (course notes here). The material I covered was adapted from Data Carpentry's Ecology lessons.
This term I'm offering a new, improved version of the same material to graduate students in CSU Biology/BSPM 526, Evolutionary Ecology. It's been really interesting experimenting with a shorter format (50 minutes over 10 weeks versus two hours over five weeks) and noting differences with learners at a more advanced stage in their education. For one, it's a lot easier to solicit questions and responses from the learners! For another, the shorter format seems to be helping me focus on what it most important to convey with each concept we cover. Or maybe practice has improved my instruction on this material. Coming up, I'll be helping out at a CSU Library Coding and Cookies basic R workshop, and teaching two half-day sessions on R at a Data Carpentry workshop as part of the National Data Integrity Conference at CSU next week. I'm excited that our paper on the genetic architecture of UV floral patterning in sunflower is the Editor's Choice for the July 2017 issue of Annals of Botany. Read it for free here!
Our second and final day of instructor training was Friday. One of the most valuable components for me was thinking about and practicing live coding. Talking through your actions as you write code allows you to troubleshoot errors as they occur, which happens all the time even to the most competent practitioners but can totally overwhelm novices. Demonstrating the actual practice of writing code helps learners develop the necessary mental models to become competent practitioners themselves.
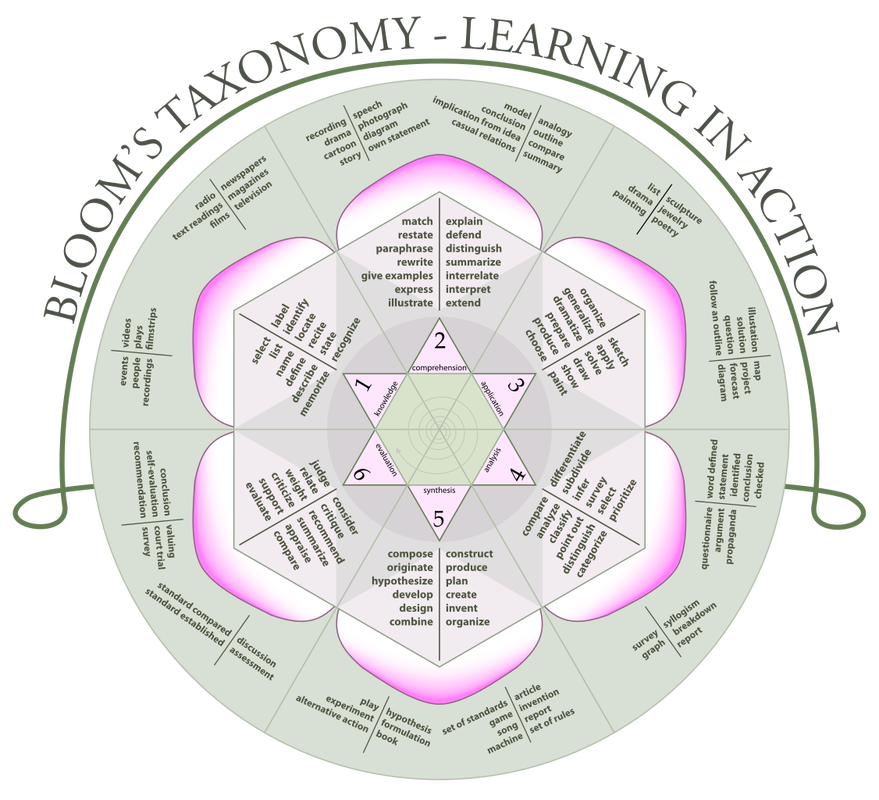
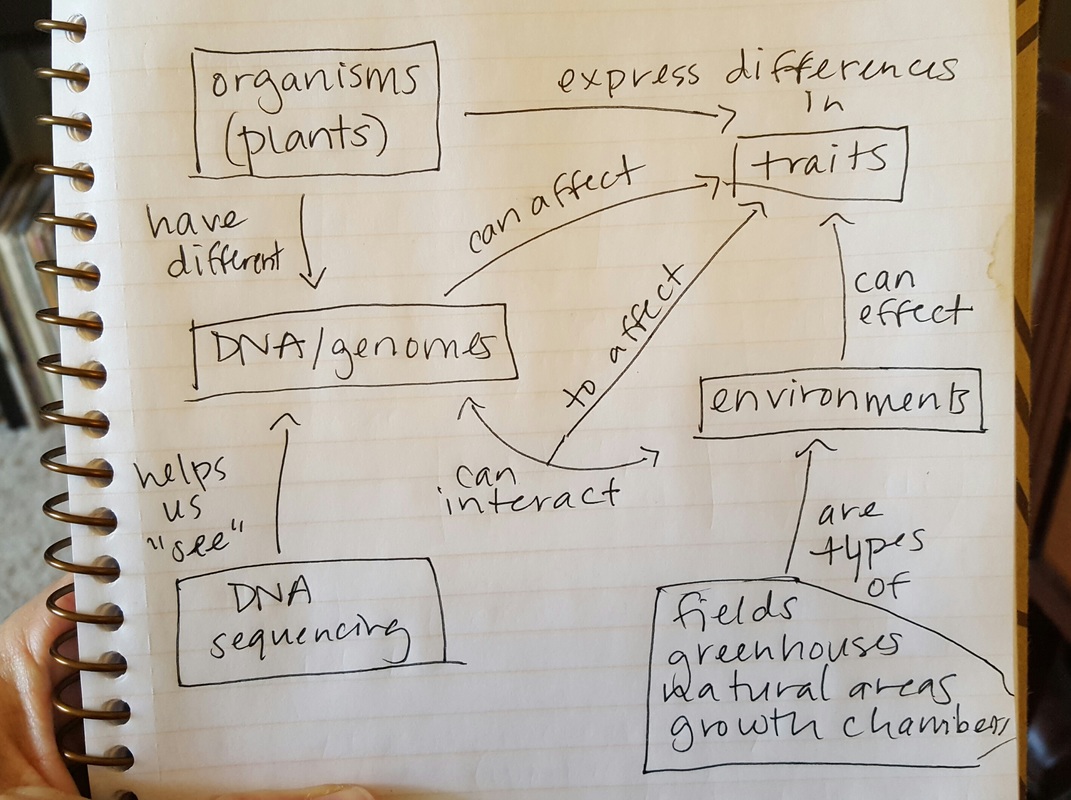
We spent a good portion of the day talking about equity in instruction, with some interesting relevant ideas for software and data carpentry (e.g. live coding isn't very helpful for visually-impaired learners). I learned that 3M brand sticky notes of different colors look different even to colorblind learners, which is useful! In these workshops, sticky notes are often used to facilitate interactions between learners and instructors, with specific colors assigned to specific flags (i.e. green: I'm following the material, red: I need assistance, yellow: I have a question, and blue: I need a break). We also talked a bit about designing lessons, starting with writing good learning objectives (using thoughtful verb choices from Bloom's taxonomy, see below), then writing summative assessments (e.g. exam questions) and rubrics, then the lessons themselves. All in all, it was a valuable experience! I look forward to teaching programming and data management more effectively in the future. I'm taking part in an instructor training for Software/Data Carpentry held online (using Blue Jeans and Etherpad, as well as additional media, and led by Greg Wilson). Day 1 involved on discussion, theory, and practice focused on educational psychology and instructional design. We practiced tiny lessons, wrote multiple choice questions with distractors, created concept maps, and developed "faded design" programming exercises. So far, I'm impressed with how much I've learned!
Important takeaways (with useful links):
Looks like I'm on an annual posting schedule here! Oh well. Last month I attended an intensive 3-week course at the International Rice Research Institute on all aspects of rice research, breeding, and production. It was a wonderful opportunity to learn about the human and global context of my research: I find it can be easy to lose sight of the bigger picture when you're down in the analysis trenches. The course also broadened my understanding of the rice breeding pipeline, which should increase the usefulness of the products of my research to breeders. Finally, I think the most valuable aspect of the course was the opportunity to interact and collaborate with scientists, extension officers, breeders, and others from all parts of the rice growing world. It was both challenging and rewarding to find common ground across so many cultures and perspectives.
I am grateful to NSF, IRRI, and co-organizers Jan Leach and Adam Bogdanove for the opportunity! |
MissionThis blog is mostly for news and occasional musings. Views belong to Brook Moyers. Some older posts mirror Brook's contributions to the Rieseberg Lab Blog. Archives
October 2021
Categories |





 RSS Feed
RSS Feed
